Decoding synaptic and cytoskeletal organization at the nanoscale
Highlights
- MINFLUX provides sub-nanometer localization precision in 2D and isotropic ∼2 nm precision in 3D
- 3D MINFLUX directly resolves synaptic protein organization at true molecular length scales
- MINFLUX resolves nanoscale protein arrangements that reveal functional insights inaccessible to diffraction-limited imaging
- MINFLUX can identify subtle and specific changes in neurological disease states
MINFLUX nanoscopy for neurobiology
Nanoscopy for neurobiology
Many of the structures and processes essential for neuronal function occur at the nanoscale. Dense synaptic protein assemblies and the periodic organization of the cytoskeleton lie well below the resolution limit of conventional light microscopy. As such, superresolution microscopy is invaluable to the neurobiology field, with stimulated emission depletion (STED), structured illumination microscopy (SIM), and camera-based single molecule localization microscopy techniques now used routinely. The resolving power of MINFLUX far exceeds older techniques, being capable of reaching sub-nanometer localization precisions in 2D and an isotropic ∼2 nm precision in 3D (1). This step change in superresolution light microscopy has been applied to a broad range of neurobiological questions, revealing structural and functional data inaccessible by other techniques.
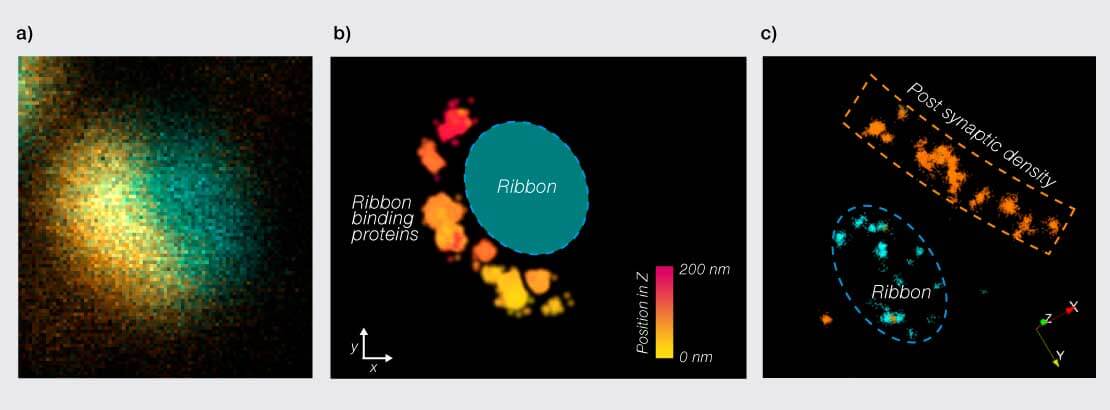
Figure 1: 3D MINFLUX imaging of inner ear hair cell ribbon synapses. a) Confocal image of a ribbon synapse with the ribbon (CtBP2/RIBEYE) stained in blue and ribbon binding protein RBP2 in orange. b) 3D MINFLUX image of the same synapse, showing the distribution of RBP2 along the active zone. Based on the confocal image, the predicted ribbon position is marked in blue. Scale bar shows relative position in the z axis. c) Two color spectral demixing MINFLUX image of another ribbon synapse, showing the ribbon (CtBP2/RIBEYE) in blue and the post synaptic density (Homer1) in orange. Data kindly provided by (2).
MINFLUX reveals nanoclusters in ribbon synapses
Studying how synapses are organized on a molecular scale can be vital to develop understanding of how they function. Sensory ribbon synapses, for example, have a highly specialized presynaptic structure for the anchoring of synaptic vesicles near the active zone. Inner ear hair cell ribbon synapses are small, crowded structures at only ∼400 nm in length. 3D MINFLUX was therefore employed in cochlear tissue sections, bridging the gap from biochemical data and electron microscopy to direct observation of single proteins forming the 3D nano-architecture of ribbon synapses (Fig.1) (2). Observation of nanoclustering behavior in Ca2+ channels and subsequent biophysical modeling suggested improved efficiency of synaptic vesicle release with this arrangement compared to random distribution, highlighting the important link between arrangement and function.
Similarly, MINFLUX may be applied to study the nano-architecture of other neuronal synapses and contribute to models of synaptic function.
Resolving the organization of the neurocytoskeleton
The membrane-associated periodic cytoskeleton (MPS) is a highly ordered periodic structure comprised of spectrin and actin, critical for neuronal integrity and regulation of inter- and intra-cellular signaling events. Macarrón-Palacios and colleagues (3) recently described the protein Palm1 as an organizer and regulator of the MPS. While STED microscopy readily resolves the characteristic ∼190 nm periodicity of Palm1 rings aligned with the actin-capping protein adducin, only MINFLUX delivers the precision required to determine their relative arrangement. 3D exchange-PAINT MINFLUX measurements revealed distances between Palm1 and adducin of <20 nm, and demonstrated that Palm1 is positioned on both sides of the adducin ring in a sandwich-like configuration.
MINFLUX could help in the same way to resolve further structural details of the neuronal cytoskeleton at the nanoscale.
Parkinson’s disease at the nanoscale
MINFLUX has significant potential for detecting nanoscale changes in neurological diseases. A recent preprint used 3D MINFLUX imaging of dopaminergic synapses in DJ-1 knockout iPSC neurons, linked to Parkinson’s disease (4). In these neurons, a global decrease in the crucial vesicular protein VMAT2 had been shown through whole cell proteomics, but the precise changes to protein distribution were unknown. Comparing the disease state to the isogenic control, MINFLUX data showed a decrease in VMAT2 at both single synapse and single vesicle levels, translating global proteomic findings into a potential nanoscale mechanism for impaired dopamine uptake.
Other neurological disorders likewise show aberrant levels as well as aggregation and mis-localization of certain proteins. MINFLUX could be used to assess open questions regarding sub-cellular localization of disease-related proteins in the nanometer range.
Conclusion
By resolving protein arrangements with nanometer precision in three dimensions, MINFLUX links structure to function, refines mechanistic models, and reveals subtle alterations in disease-relevant systems. This makes MINFLUX a uniquely powerful platform for advanced neurobiological research at the molecular level, able to provide nanoscale information that is inaccessible with other techniques.
References
- Schmidt R, Weihs T, Wurm CA, Jansen I, Rehman J, et al. MINFLUX nanometer-scale 3D imaging and microsecond-range tracking on a common fluorescence microscope. Nat Commun. 12, 1478 (2021). doi: 10.1038/s41467-021-21652-z
- Kapoor R, Kim H, Garlick E, Lima MAdRBF, Esch K, et al. Charting the nanotopography of inner hair cell synapses using MINFLUX nanoscopy. Sci Adv. 11, eady4344 (2025). doi:10.1126/sciadv.ady4344
- Macarrón-Palacios V, Hubrich J, Lima MAdRBF, et al. Paralemmin-1 controls the nanoarchitecture of the neuronal submembrane cytoskeleton. Sci Adv. 11, eadt3724 (2025). doi: 10.1126/sciadv.adt3724
- Heger LM, Gubinelli F, Huber A, Cardona-Alberich A, et al. LF. VMAT2 dysfunction impairs vesicular dopamine uptake, driving its oxidation and α-synuclein pathology in DJ-1-linked Parkinson’s disease neurons. bioRXiv. 2025.09.22.677565 (2025). doi: 10.1101/2025.09.22.677565
Author
Evelyn Garlick




